MS-FINDER tutorial
Last edited in 2015/010/19
Abstract
The purpose of metabolomics is to perform the ‘comprehensive’ analysis for small biomolecules of
living organisms. Liquid chromatography coupled with electrospray ionization- (ESI-) tandem mass
spectrometer (LC/MS/MS) is the preferred tool for untargeted metabolomics. Currently, the main
bottleneck of LC/MS/MS based untargeted analysis is compound identification due to the limitation
of MS/MS records of authentic standard.
MS-FINDER aims to provide solutions for 1) formula predictions, 2) fragment annotations,
and 3) structure elucidations by means of survey scan MS and MS/MS spectra of unknowns. MS-
FINDER has been developed as the collaborative work between Prof. Masanori Arita team (RIKEN,
Reifycs Inc.) and Prof. Oliver Fiehn team (UC Davis) supported by the JST/NSF SICORP
“Metabolomics for the low carbon society” project.
Hiroshi Tsugawa
RIKEN Center for Sustainable Resource Science
hiroshi.tsugawa@riken.jp
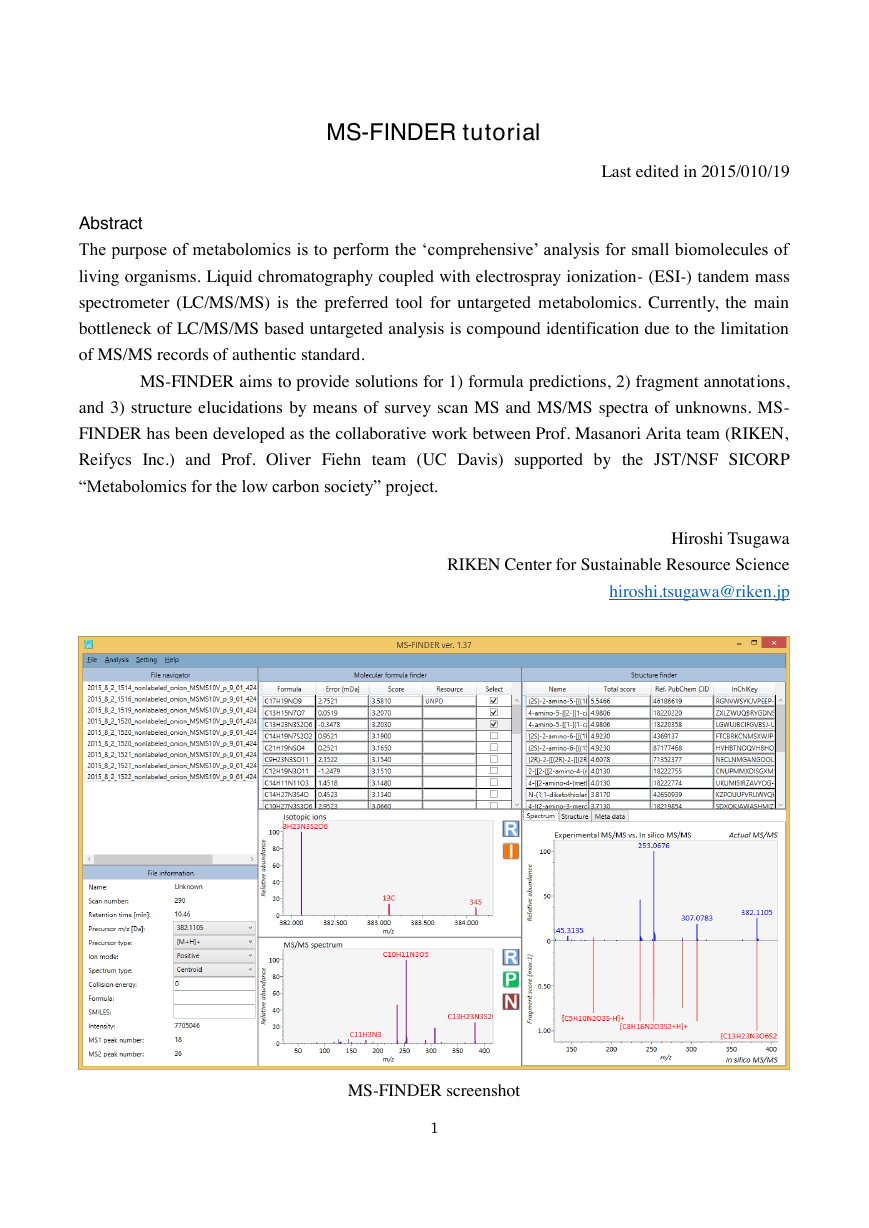
MS-FINDER screenshot
1
�
2
�
Table of contents
Software environments .......................................................................................................................... 4
Required programs ................................................................................................................................ 5
Acceptable ASCII formats ...................................................................................................................... 6
MSP format ........................................................................................................................................ 6
MAT format ........................................................................................................................................ 7
Adduct ion format: [M+Na]+, [M+2H]2+, [M-2H2O+H]+, [2M+FA-H]-, etc. ..................................... 9
Import queries ..................................................................................................................................... 10
A. From a folder which includes MSP or MAT format files ............................................................. 10
B. From the graphical user interface of the MS-FINDER program ................................................ 11
C. From the MS-DIAL program ....................................................................................................... 12
Analysis parameter setting ................................................................................................................. 13
Formula finder tab ........................................................................................................................... 13
Structure finder tab ......................................................................................................................... 15
Compound annotation (Single analysis) .............................................................................................. 17
Compound annotation (batch analysis) ............................................................................................... 19
Peak assignment (single) ..................................................................................................................... 20
Peak assignment (batch job) ................................................................................................................ 21
Mouse function ..................................................................................................................................... 22
Export .................................................................................................................................................. 23
Help ...................................................................................................................................................... 24
3
�
Software environments
Windows OS (.NET Framework 4.0 or later): Windows 7 or later
RAM: 8.0 GB or more
4
�
Required programs
MS-FINDER
Download link: http://prime.psc.riken.jp/Metabolomics_Software/MS-FINDER/index.html
MS-FINDER can be used as the local software program in Windows PC. The program can
import ASCII format files including MSP (MS/MS) or improved MSP (both MS and MS/MS,
the file extension should be MAT.). In addition, the users can directly make the query in the MS-
FINDER graphical user interface. Moreover, this program can be called from the MS-DIAL
program which is downloadable at http://prime.psc.riken.jp/Metabolomics_Software/MS-
DIAL/index.html.
5
�
Acceptable ASCII formats
This program accepts two file extensions, i.e. MSP or MAT formatted by the following explanations.
The point is that unknown queries should be separately stored in the ASCII file: the MSP or MAP
file ‘CANNOT’ store multi compound records in the single file.
MSP format
The format of MSP basically follows the NIST MS search manual.
Link: http://www.nist.gov/srd/upload/NIST1a11Ver2-0Man.pdf
Required fields
NAME:
PRECURSORMZ:
PRECURSORTYPE:
IONMODE: (Positive or Negative)
SPECTRUMTYPE: (Centroid or Profile)
Num Peaks:
m/z intensity pair (tab, comma, space can be used as the delimiter.)
Mass spectrum is supposed to be imported as MS/MS.
If you want to perform the MS/MS peak annotation with the known structure, prepare two fields
including FORMULA and SMILES. Please note that the formula and SMILES of the neutralized
structure should be prepared.
MSP example
6
�
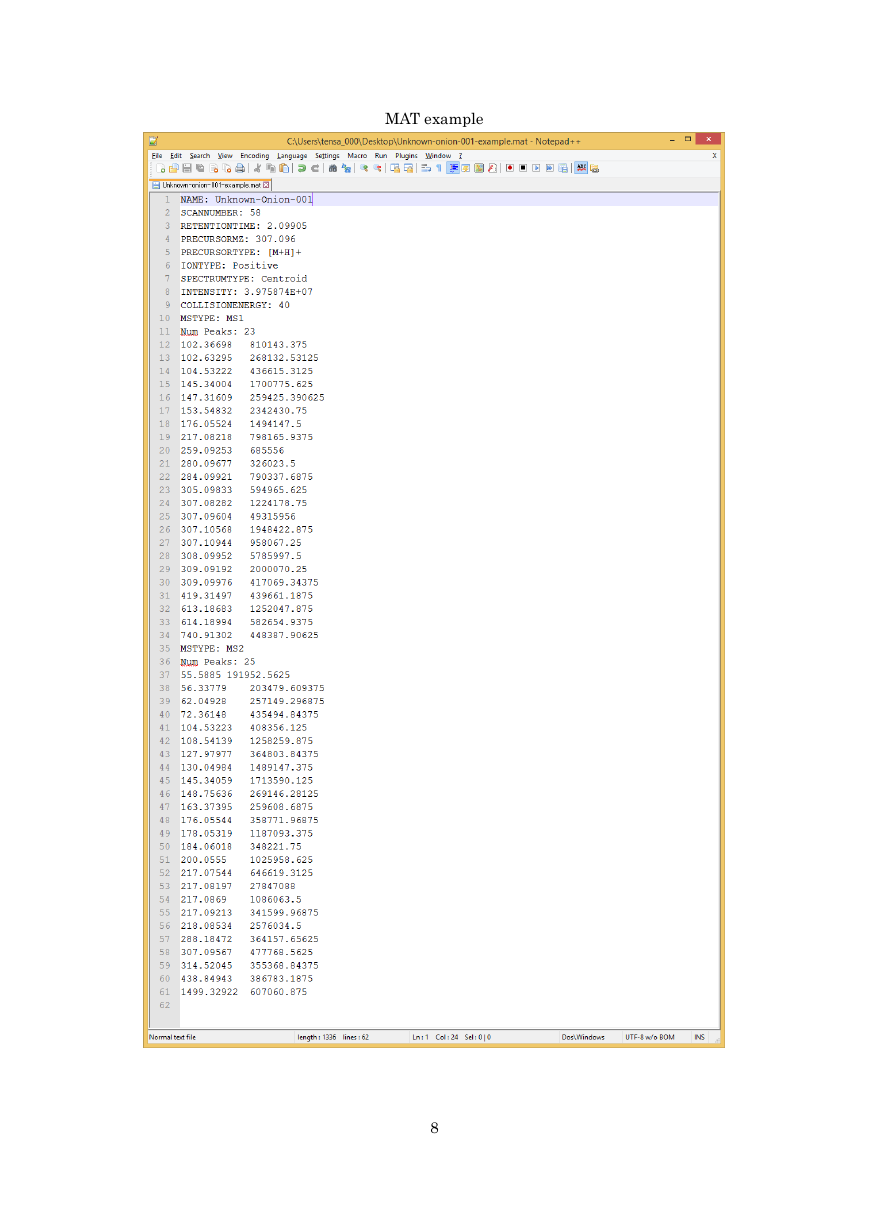
MAT format
The MAT format was defined as the improved version of MSP in the MS-FIDNER program to store
both MS1 and MS/MS spectra in the same file. The survey scan MS data should be required to
calculate ‘isotopic ion score’ for formula predictions.
Required fields
NAME:
PRECURSORMZ:
PRECURSORTYPE:
IONMODE:
MSTYPE:
Num Peaks:
m/z intensity pair (tab, comma, space can be used as the delimiter.)
Three fields including MSTYPE, Num Peaks, and m/z intensity pair should be SERIALLY
stored.
If you type ‘MSTYPE: MS1’, the spectrum written from next field should be recognized as the
survey scan MS (MS1). If you type ‘MSTYPE: MS2’, next spectrum should be recognized as
the MS/MS spectrum.
Both field (MSTYPE: MS1 and MSTYPE: MS2) is not necessary for this program, i.e. the users
can import the ASCII file as only MS1 spectrum or as only MS/MS spectrum record.
Actually, the users may prepare the MAT or MSP files without any spectrum record. In such
case, the formula prediction will be performed by means of mass accuracy and database criteria.
If you want to perform the MS/MS peak annotation with the known structure, prepare two fields
including FORMULA and SMILES. The formula and SMILES of the neutralized structure
should be made.
7
�
MAT example
8
�















 2023年江西萍乡中考道德与法治真题及答案.doc
2023年江西萍乡中考道德与法治真题及答案.doc 2012年重庆南川中考生物真题及答案.doc
2012年重庆南川中考生物真题及答案.doc 2013年江西师范大学地理学综合及文艺理论基础考研真题.doc
2013年江西师范大学地理学综合及文艺理论基础考研真题.doc 2020年四川甘孜小升初语文真题及答案I卷.doc
2020年四川甘孜小升初语文真题及答案I卷.doc 2020年注册岩土工程师专业基础考试真题及答案.doc
2020年注册岩土工程师专业基础考试真题及答案.doc 2023-2024学年福建省厦门市九年级上学期数学月考试题及答案.doc
2023-2024学年福建省厦门市九年级上学期数学月考试题及答案.doc 2021-2022学年辽宁省沈阳市大东区九年级上学期语文期末试题及答案.doc
2021-2022学年辽宁省沈阳市大东区九年级上学期语文期末试题及答案.doc 2022-2023学年北京东城区初三第一学期物理期末试卷及答案.doc
2022-2023学年北京东城区初三第一学期物理期末试卷及答案.doc 2018上半年江西教师资格初中地理学科知识与教学能力真题及答案.doc
2018上半年江西教师资格初中地理学科知识与教学能力真题及答案.doc 2012年河北国家公务员申论考试真题及答案-省级.doc
2012年河北国家公务员申论考试真题及答案-省级.doc 2020-2021学年江苏省扬州市江都区邵樊片九年级上学期数学第一次质量检测试题及答案.doc
2020-2021学年江苏省扬州市江都区邵樊片九年级上学期数学第一次质量检测试题及答案.doc 2022下半年黑龙江教师资格证中学综合素质真题及答案.doc
2022下半年黑龙江教师资格证中学综合素质真题及答案.doc